|
3/23/2021 0 Comments Molecular Docking Tools
MS supports SDF, MOL, MOL2, PDB and all other common file formats.You can SAVE the ligand with correct protonation state in your desired file format.
 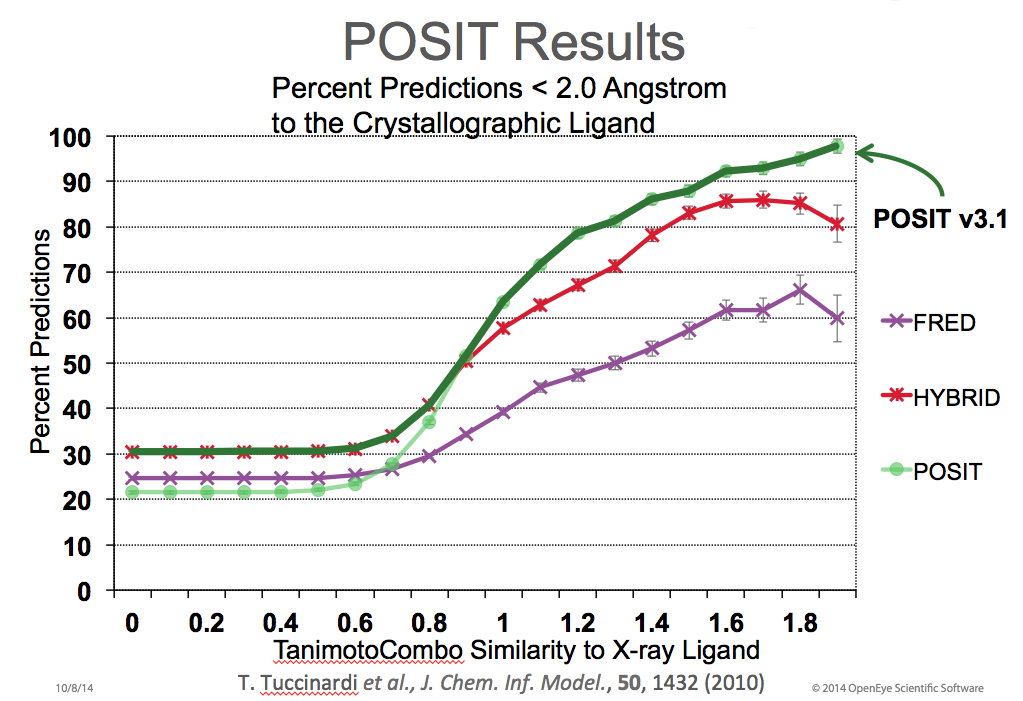
You can upload the PDB structure (remove waters and other non-standard residues before) and upload. You can see it also gives dimension of Center X, Y and Z corresponds to each binding site which you can use as Autodock Vina input (in the VS.bash file, scroll down and follow the tutorial). You can click on each cluster and see the residues correspond to that cluster. You can see that the non-terminal missing residues are highlighted as dot marks. Molecular Docking Tools License Key Which YouCheck the boxes as follows: Modelremodel: non-terminal missing structure Allow this many residues adjacent to missing regions to move: 1 Number of models to generate: 5 Loop modelling protocol: standard Run Modeller using: Choose either web service or local installation Modeller license key: Put the modeller license key which you can get from MODELLER website. Clicking the information icon near the bottom of the Chimera window will bring up the Task Panel, in which the job can be canceled if desired. The models can be viewed individually or collectively by choosing rows in the dialog with the mouse. The different scores from Modeller use different criteria and will not necessarily agree on which models are best. Displaying all the models at once shows little conformational variability except in the termini, and to a lesser extent, the untemplated part of the third intracellular loop. This conclusion is reinforced by the RMSD histogram in the sequence window, where bar heights indicate root-mean-square distances among the -carbons of the residues associated with a column. You can see the installation instruction in the Autodock Vina webpage. In my point of view testing all the docking softwares and protocols in the blind prediction challenge will be the ultimate test for all the docking softwares.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
